文章轉載自中央研究院-院新聞稿
自閉症譜系障礙(autism spectrum disorder,簡稱ASD)是一種腦部發育障礙所導致的複雜疾病,患者往往在社交溝通、互動及表達上有障礙,成因目前仍未有定論,普遍認為與遺傳及基因變異有關。中央研究院基因體研究中心研究員莊樹諄研究團隊,首次系統性建構環狀RNA(circular RNA1)在自閉症腦部的基因調控網路圖譜,有助於增進對自閉症致病分子機制的理解。該篇論文已於今(109)年3月刊登在《基因體研究》(Genome Research)。
環狀RNA是一種單鏈封閉式環型結構,且特別高度表現在神經系統。莊樹諄研究團隊利用大數據分析找到在自閉症患者大腦皮質中表現量異常的環狀RNA,並預測其調控路徑,結合分子生物實驗後證實:環狀RNA像海綿一樣吸附特定的微RNA(miRNA),使其失去或降低對下游自閉症風險基因調控的能力。有關環狀RNA、微RNA、與下游基因在自閉症腦部的調控網路關係,過去並未被有系統地探討。
莊樹諄所率領的大數據分析與神經科學實驗室團隊,透過先前開發的環狀RNA偵測軟體(NCLscan),設計大數據分析流程。從超過200個樣本的轉錄體定序(RNA-seq)資料,找到60個在自閉症患者大腦皮質中表現異常的環狀RNA;經統計模型分析顯示,根據此60個環狀RNA的表現情形,能有效區別自閉症與非自閉症樣本,因此可判定這些環狀RNA與自閉症的發生應有關連 (圖一)。
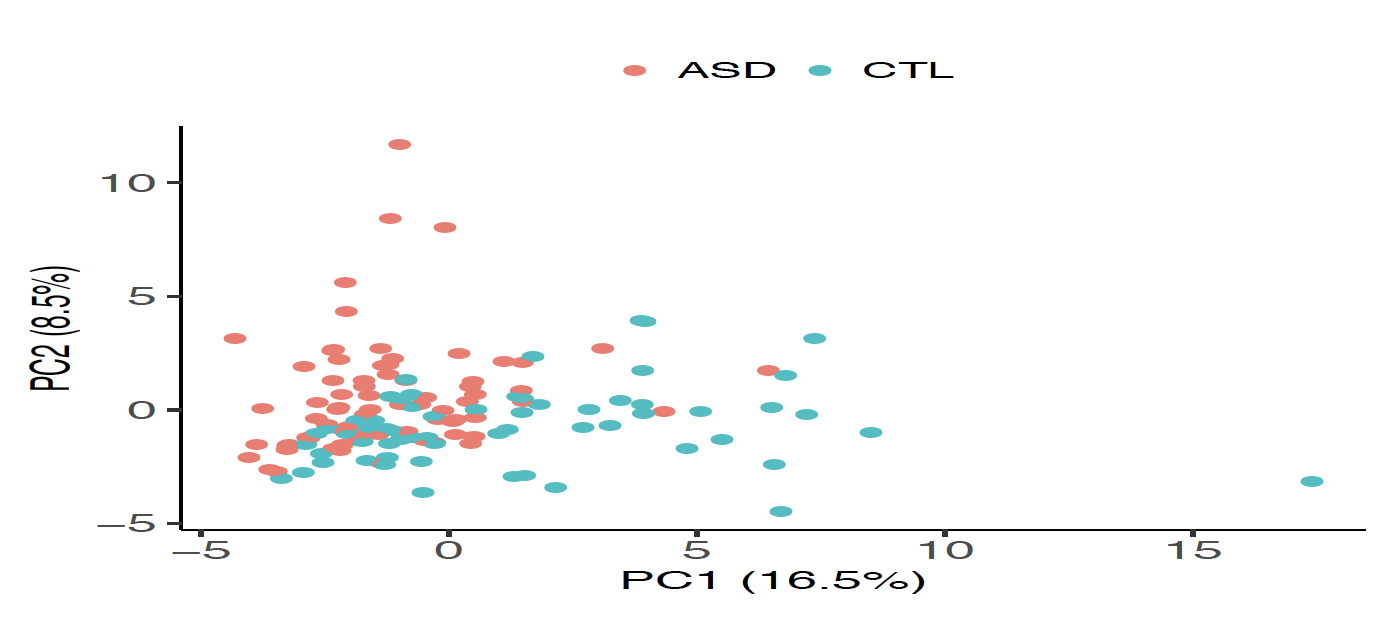 |
| 圖一、偵測在自閉症患者大腦皮質中表現異常的60個環狀RNA。A圖顯示22個(紅點)表現量在自閉症患者顯著上升, 38個(綠點)表現量在自閉症患者顯著下降,其餘(灰點)表示在自閉症與非自閉症者間無顯著差異。在此每一點表示一個環狀RNA。B圖顯示這60個環狀RNA的表現量能有效區別自閉症(紅點)和非自閉症(綠點)樣本。在此每一點表示一個腦組織樣本。 |
環狀RNA調控網路和自閉症風險基因高度相關
為此,團隊進一步預測這些環狀RNA的下游調控路徑,建構出8,170個環狀RNA、微RNA、信使RNA(mRNA)2間的交互調控網路 (圖二),接著再透過基因富集分析3,發現這些網路所調控的下游目標基因,顯著集中在已知的自閉症風險基因。
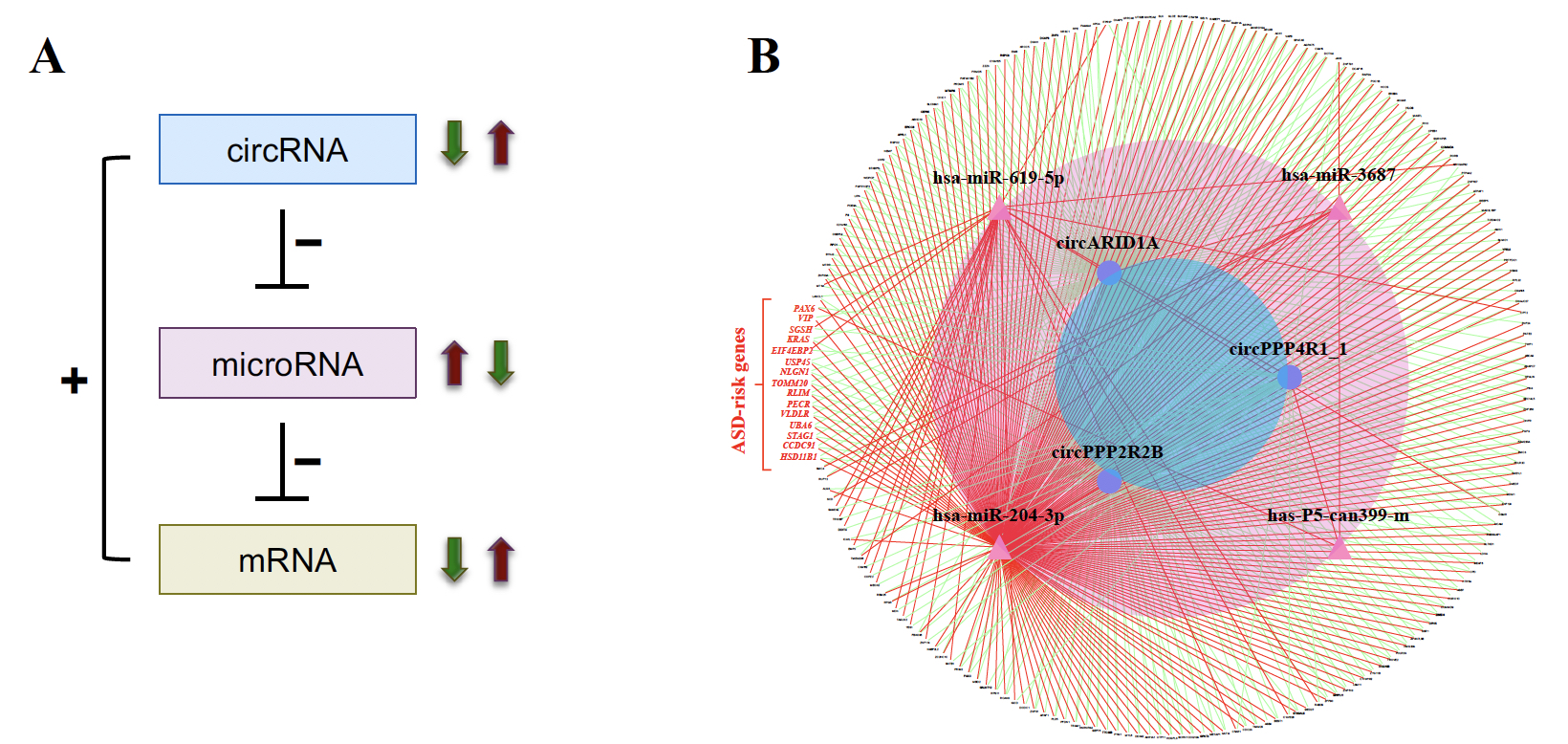 |
| 圖二、環狀RNA調控網路。A圖為環狀RNA、微RNA、信使RNA間交互調控網路示意圖。B圖為所預測的其中部分的調控網路,紅色字體顯示此網路中的12個已知的自閉症風險基因。 |
莊樹諄說明,這個研究除設計大數據分析流程來建構環狀RNA的調控網路關係,也結合分生實驗驗證。團隊挑選一個在自閉症患者腦部表現量明顯上升的環狀RNA(命名為circARID1A),於人類神經細胞實驗驗證後發現, circARID1A確實可藉由調控微RNA(miR-204-3p),影響下游多個自閉症風險基因的表達 (圖三)。
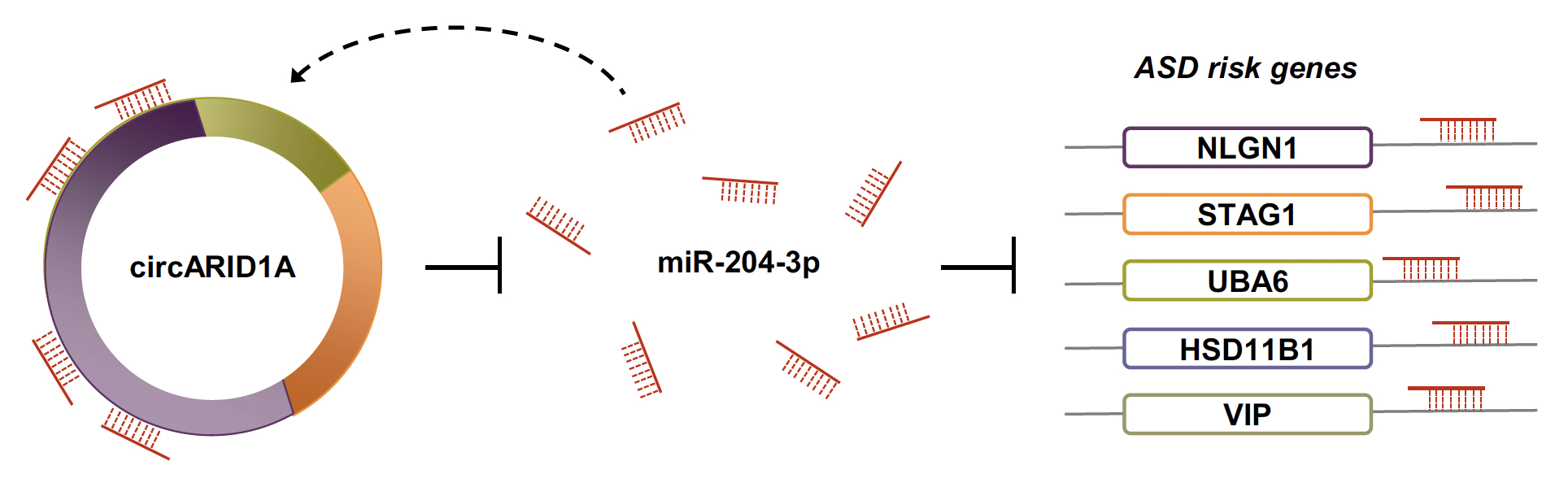 |
| 圖三、在人類神經相關細胞(NHA或ReN cells)實驗驗證circARID1A確實可藉由調控miR-204-3p影響自閉症風險基因(如NLGN1、 STAG1、HSD11B1、VIP、UBA6)的基因表達。 |
論文封面圖片靈感出自紀錄片《遙遠星球的孩子》
莊樹諄團隊不僅建構環狀RNA的調控網路,論文封面也讓人好奇,只見一個孩子孤獨地待在自己的星球上,寂寞地望著地球。
 |
| 此封面圖片設計靈感來自於紀錄片《遙遠星球的孩子》,因無法融入地球軌道,一個人孤獨地待在自己的星球上。繪圖/徐維駿 |
莊樹諄解釋,此圖片設計靈感來自於以自閉症為主題的紀錄片《遙遠星球的孩子》,環繞在星球外圍的光環即環狀RNA,像海綿一樣吸附軌道上的小行星,「像上帝畫的圈圈」,讓自閉症孩子只能待在自己的星球上,難以融入地球常軌。
由於自閉症發生原因不明,本文揭開環狀RNA在自閉症腦組織的調控關係,可用以探究自閉症的致病分子機制,對於未來診斷、追蹤及治療提供新的思考方向。研究團隊結合資訊、統計、分生、演化等知識背景 (圖四),所設計的大數據分析與分生實驗流程,將來也可應用於與環狀RNA調控相關之其他神經疾病上,如:阿茲海默症、帕金森氏症、思覺失調症等。
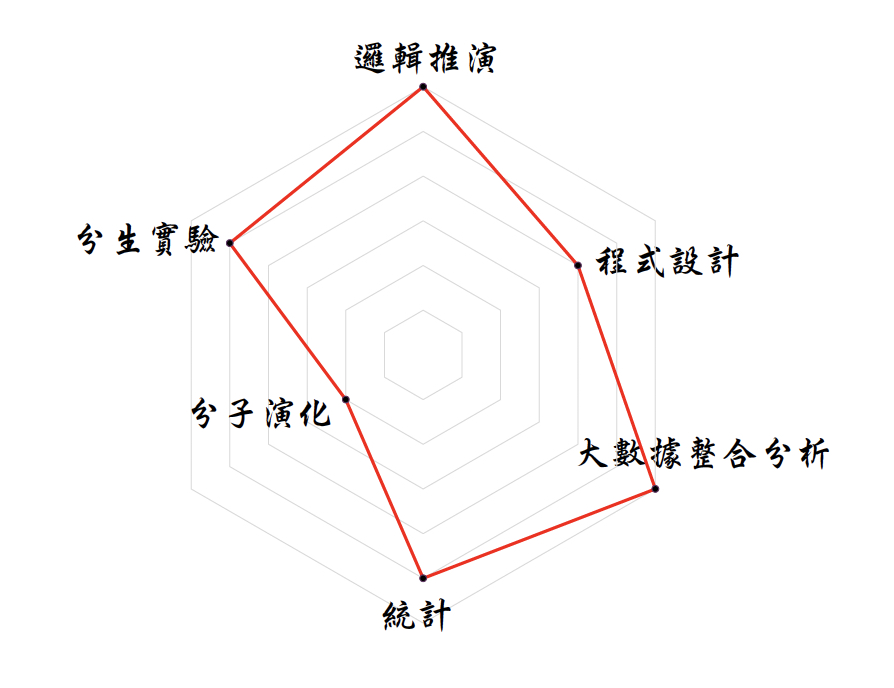 |
| 圖四、本研究所需的知識背景雷達圖。目前該研究團隊中有一半是「純理工」背景,另一半是「純分子生物」背景,一個理想的生物資訊團隊極需要上述兩種背景的同仁緊密合作。歡迎具備上述任一背景的同學加入嶄新的醫療大數據分析世界。詳情請上莊樹諄老師實驗室網站查詢:http://idv.sinica.edu.tw/trees/ |
本論文共同第一作者包括基因體研究中心陳彥如、陳嘉瑩、麥德倫博士,通訊作者為莊樹諄研究員。
全文詳見:https://genome.cshlp.org/content/30/3/375.full.pdf+html
註釋:
1. 環狀RNA因其構型關係不易被核酸外切酶降解,因此比其他RNA更加穩定,適合開發為新型臨床診斷的生物標記。近年來隨著次世代定序技術快速發展,陸續發現大量的環狀RNA存在各種生物體中。
2. miRNA會抑制mRNA的基因表達,當miRNA結合至目標基因mRNA序列上,使mRNA無法進行轉譯作用而產生蛋白質。當環狀RNA吸附miRNA,將使miRNA失去或降低其調控下游基因表達的能力。
3. 基因富集分析(gene set enrichment analysis) 是一種統計分析策略,用以針對某一組特定基因,探討這組基因是否特別表現在哪一種(或多種)已知功能。