Introduction
Currently,“peptide synthesis” includes a large range of techniques and procedures that enable the preparation of materials ranging from small peptides to large proteins. The pioneering work of Bruce Merrifield, which introduced solid phase peptide synthesis (SPPS), dramatically changed the strategy of peptide synthesis and simplified the tedious and demanding steps of purification associated with solution phase synthesis. Moreover, Merrifield’s SPPS also permitted the development of automation and the extensive range of robotic instrumentation now available. After defining a synthesis strategy and programming the amino acid sequence of peptides, machines can automatically perform all the synthesis steps required to prepare multiple peptide samples. SPPS has now become the method of choice to produce peptides, although solution phase synthesis can still be useful for large-scale production of a given peptide.
This core facility provides synthetic peptides to support the research programs in GRC. Peptides with various modifications (phosphorylation, glycosylation, methylation etc.) or with fluorescence labeling can also be provided. Under the collaboration with other research groups in GRC, we developed the methods for synthesizing several newly discovered peptides and their derivatives which can not be prepared by routine methods.
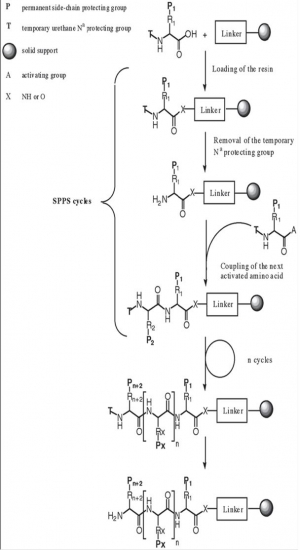
Principles of SPPS
Solid Phase peptide synthesis (SPPS) is based on sequential addition of alpha-amino and side-chain protected amino acid residues to an insoluble polymeric support .
The acide-labile Boc group or base-labile Fmoc-group is used for N-alpha-protection. After removal of this protecting group, the next protected amino acid is added using either a coupling reagent or pre-activated protected amino acid derivative. The resulting peptide is attached to the resin, via a linker, through its C-terminus and may be cleaved to yield a peptide acide or amide, depending on the linking agent used. Side chain protecting groups are often chosen so as to be cleaved simultaneously with detachment of the peptide from the resin.
Equipments and Materials
Peptide Purification
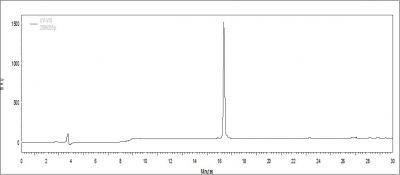
The purity of a peptide is often the spokesman of the wheel that dictates the results of future proceeding experiments. Peptide length, complexity, and order as well as side modification are all immediate factors that can downgrade its purity. To ensure all of our fellow researchers in GRC to conduct experiment smoothly, our laboratory uses the new HPLC model from Aglient 1200 and Hitachi L-2420, which guarantees most peptides to have > 95% in purity. Below is a HPLC elution profile of a purified compound. The peak represents the peptide desired.
Peptide Confirmation
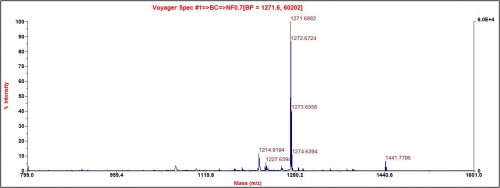
Our peptides are confirmed by using Mass Spectrometry (MALDI). This ensures the molecular weight of the actual peptide is consistent with the theoretical weight. Below is a mass profile of confirmed Peptide with a molecular weight of 1272.
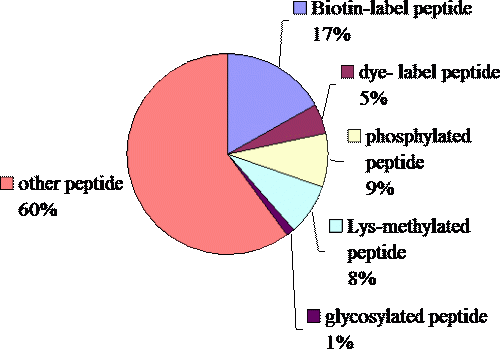
The variety of the peptides synthesized in this core facility. The group “other” includes cyclized peptide, peptide with disulfide bridge, N-acetylation, C-amidated peptide, D-form peptide, Aβ40,42 analog peptide (5-43 residue), TPA labeled peptide, cycl-peptide, Lysine-branch-peptide etc..
Various Peptide Applications
Biotinylation, phosphorylation (tyrosine, serine, threonine), Amide, Acetylation, hydroxylation, Fluorochrome, Sulfation, Nitration, and cyclization etc., are all avaliable in our lab. Here are some examples.
A. Kinase Bind Assay
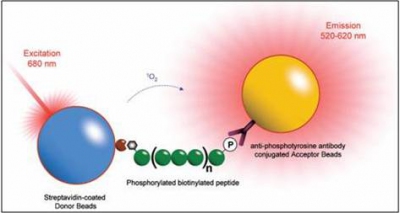
A sequence of a phosphorylated biotinylated peptide is often used to detect whether a sample protein is biologically active to the specific peptide sequence. The picture describes how the phosphorylated peptide triggers the steptavidin-coated donor beads (attached to biotin in red) to excite the phosphate binding antibody to responsd.
B. Antibody inducing peptides
Custom Peptides can be designed and synthesized to induce a specific antibody.
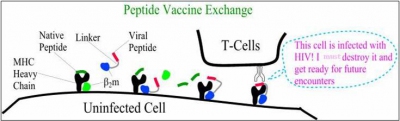
C. Peptide Vaccine
Recently the Institute of Plant and Microbial center here at Academia Sinica have developed a recombinant bamboo mosaic virus (BaMV) expressing a the antigenic epitope(s)(peptide) of a specific protein as a vaccine vector. Similar strategies have also been employed in HIV and cancer vaccines studies