 Professor
Professor
cma
+886-2-27871233
+886-2-27871233
EDUCATION AND POSITIONS HELD:
- B.S., Chemistry, National Taiwan University, 1992
- M.S., Chemistry, University of Pennsylvania, 1996
- Ph.D., Chemistry, University of Pennsylvania, 2000
- Postdoctoral Fellow, University of California at San Diego, 2001
- Postdoctoral Fellow, The Scripps Research Institute, 2001–2004
- Assistant Professor, Genomics Research Center, Academia Sinica, Taiwan, 2004-2010
- Associate Professor, Genomics Research Center, Academia Sinica, Taiwan, 2010-2019
- Visiting Scientist, RIKEN SPring-8 Center, Japan, 2016 – present
- Division Director of Chemical Biology, GRC, Academia Sinica, Taiwan, 2016 – 2023
- Professor, Genomics Research Center, Academia Sinica, Taiwan, 2019-present
HONORS:
- University of Pennsylvania, Teaching Assistant Award, 1995
- Keystone Symposia Scholarship, Frontiers of Structural Biology, 2003
- The Skaggs Postdoctoral Fellowship, 2001–2003
- TWAS Young Affiliate, 2009-2013
- Academia Sinica Significant Research Achievements, 2009
- Academia Sinica Research Award for Junior Research Investigators, 2010
- Two research highlights in Taiwan Yearbook of Technology, 2010
- The Young Scholar Award of Tien-De Li Biomedical Foundation, 2011
- Academia Sinica Significant Research, 2012
- Academia Sinica Career Development Award, 2013
- Exceptional Merit in Academic Award from Chung Hwa Rotary Educational Foundation, 2014
- Taiwan Bio-Development Foundation Chair in Biotechnology, 2014
- Two FUTEX Futuretech Awards, Ministry of Science and Technology, Taiwan
RESEARCH INTERESTS:
Structure and functions of membrane glycoproteins in drug discovery
- In order to overcome the problems of drug-resistant bacterial infection, the membrane-bound bifunctional transglycosylase, has been chosen for structural and functional analysis. We have determined the X-ray crystal structures of this membrane-bound enzyme in complex with its inhibitor moenomycin, and with its substrate lipid II analog, leading to the proposed mechanism of peptidoglycan synthesis. In addition, the crystal structure of a membrane protein complex, the key component of divisome ftsBLQ has been determined to decipher the cell division process.
- Targeting the bacterial transglycosylase: antibiotic development from a structural perspective, X Chen, CH Wong, C Ma*. ACS Infectious Diseases, 5, 1493 (2019). (Times cited: 16)
- Structure of the heterotrimeric membrane protein complex FtsB-FtsL-FtsQ of the bacterial divisome. HTV Nguyen, X Chen, C Parada, AC Luo, O Shih, US Jeng, CY Huang, YL Shih* and C Ma*. Nat Commun. 14(1):1903 (2023).
- We have discovered a simple and practical approach using traditional egg-based method to prepare a broad-spectrum split influenza virus vaccine that when vaccinated can induce cross-strain protection in mice.
- Egg-based influenza split virus vaccine with monoglycosylation induces cross-strain protection against influenza virus infections, YC Tseng, CY Wu, ML Liu, TH Chen, WL Chiang, YH Yu, JT Jan, KI Lin, CH Wong*, C Ma*. Proc Natl Acad Sci U S A, 116, 4200 (2019). (Times cited: 28)
- Better influenza vaccines: an industry perspective. JR Chen, YM Liu, YC Tseng, C Ma*. J Biomed. Sci., 27, 33 (2020). (Times cited: 66)
- Chimeric hemagglutinin vaccine elicits broadly protective CD4 and CD8 T cell responses against multiple influenza strains and subtypes. HY Liao, SC Wang, YA Ko, KI Lin, C Ma, TJR Cheng, CH Wong. Proc Natl Acad Sci U S A, 117, 17757 (2020). (Times cited: 18)
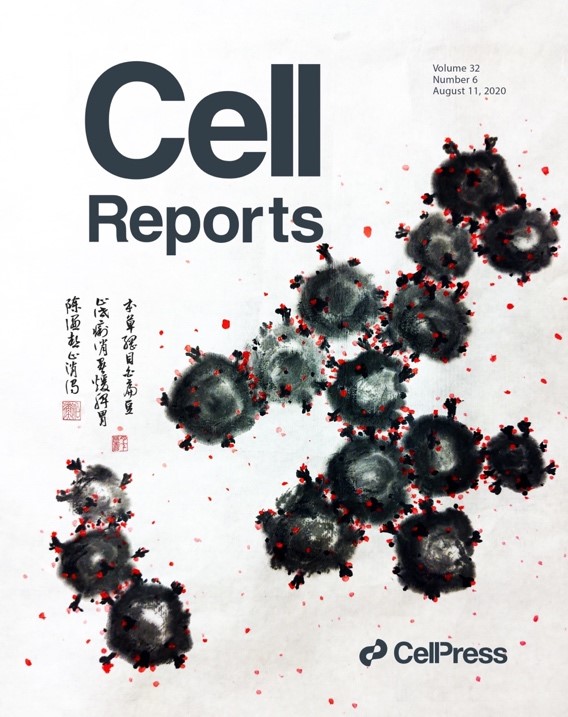
- We isolated a plant lectin FRIL from the edible Lablab beans with superior anti-influenza and anti-SARS-CoV-2 activities. FRIL binds preferentially to complex-type N-glycans and neutralizes viruses that possess complex-type N-glycans on their envelopes. As a homotetramer, FRIL is capable of aggregating influenza particles through multivalent binding and trapping influenza virions in cytoplasmic late endosomes, preventing their nuclear entry. Remarkably, FRIL also effectively neutralizes SARS-CoV-2, preventing viral protein production and cytopathic effect in host cells. These findings suggest a potential application of FRIL for the prevention and/or treatment of influenza and COVID-19.
- A carbohydrate-binding protein from the edible Lablab beans effectively blocks the infections of influenza viruses and SARS-CoV-2. YM Liu, M. Shahed-Al-Mahmud, X Chen, TH Chen, KS Liao, JM Lo, YM Wu, MC Ho, CY Wu, CH Wong, JT Jan, C Ma*. Cell Reports, 32, 108016 (2020), selected as journal cover. (Times cited: 62)
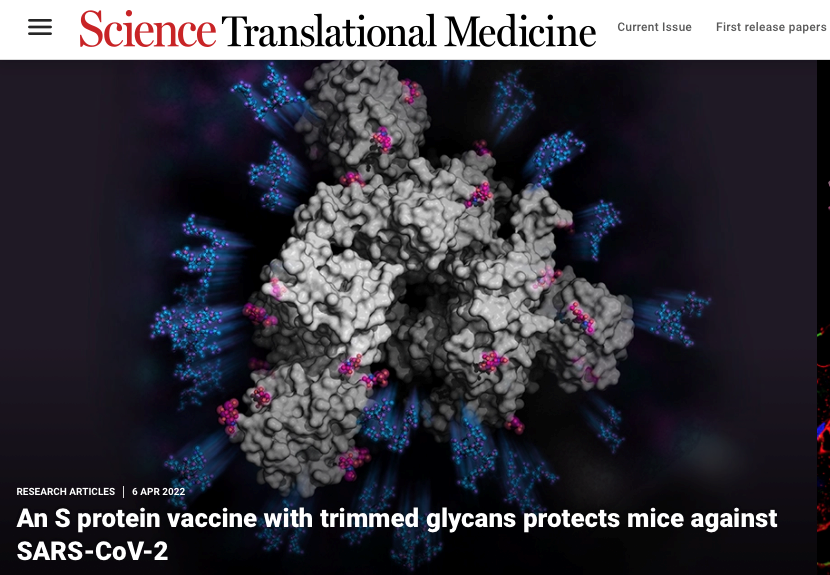
- We have studied the effect of glycosylation on SARS-CoV-2 virus major surface Spike protein with regards to its role in ACE2 receptor binding and immune response, and developed a simple strategy for broad-spectrum COVID-19 vaccine design. The resulting mono-GlcNAc-decorated Spike vaccination can elicit superior B cell, Tfh cell and CD8 T cell responses, and protect hACE2 transgenic mice against infections of SARS-CoV-2 and the variants of concerns, such as the original Wuhan, Alpha, Beta, Gamma, Delta and more recently Omicron and its subvariants. Antibody responses from a structural perspective are also been studied with samples from COVID-19 vaccination and convalescent individuals. Several structures have been determined to understand the structural basis of anti-RBD neutralizing antibodies.
- Vaccination with SARS-CoV-2 spike protein lacking glycan shields elicits enhanced protective responses in animal models. HY Huang, HY Liao, X Chen, SW Wang, CW Cheng, M Shahed-Al-Mahmud, YM Liu, A Mohapatra, TH Chen, JM Lo, YM Wu, HH Ma, YH Chang, HY Tsai, YC Chou, YP Hsueh, CY Tsai, PY Huang, SY Chang, TL Chao, HC Kao, YM Tsai, YH Chen, CY Wu, JT Jan, TJR Cheng, KI Lin*, C Ma* and CH Wong*. Science Translational Medicine, 14, eabm0899 (2022), selected as online feature article. (Times cited: 33)
- Glycosite-deleted mRNA of SARS-CoV-2 spike protein as a broad-spectrum vaccine. CY Wu, CW Cheng, CC Kung, KS Liao, JT Jan, C Ma and CH Wong*. Proc Natl Acad Sci U S A, 119, e2019995119 (2022). (Times cited: 13)
- Structures and therapeutic potential of anti-RBD human monoclonal antibodies against SARS-CoV-2. KYA Huang, D Zhou, TK Tan, C Chen, HME Duyvesteyn, Y Zhao, HM Ginn, L Qin, P Rijal, L Schimanski, R Donat, A Harding, J Gilbert-Jaramillo, W James, JA Tree, K Buttigieg, M Carroll, S Charlton, CE Lien, MY Lin, CP Chen, SH Cheng, X Chen, TY Lin, EE Fry, J Ren, C Ma, AR Townsend, DI Stuart. Theranostics, 12, 1 (2022). (Times cited: 5)
- Breadth and function of antibody response to acute SARS-CoV-2 infection in humans. KYA Huang, TK Tan, TH Chen, CG Huang, R Harvey, S Hussain, CP Chen, A Harding, J Gilbert-Jaramillo, X Liu, M Knight, L Schimanski, SR Shih, YC Lin, CY Cheng, SH Cheng, YC Huang, TY Lin, JT Jan, C Ma, W James, RS Daniels, Rodney JW McCauley, P Rijal, Pramila, AR Townsend. PLOS Pathogens, 17, e1009352 (2021). (Times cited: 55)
- Structural basis for a conserved neutralization epitope on the receptor-binding domain of SARS-CoV-2. KA Huang*, X Chen, A Mohapatra, HTV Nguyen, L Schimanski, TK Tan, P Rijal, SK Vester, RA Hills, M Howarth, JR Keeffe, AA Cohen, LM Kakutani, YM Wu, M Shahed-Al-Mahmud, YC Chou, PJ Bjorkman, AR Townsend, C Ma*. Nat Commun. 14(1):311 (2023).
- We have generated a monoclonal antibody to target an important membrane receptor IL-17RB for pancreatic cancer, which has an extremely high mortality rate due to its aggressive metastatic nature. Resolving the underlying mechanisms will be crucial for treatment. We found that overexpression of IL-17B receptor (IL-17RB) strongly correlated with postoperative metastasis and inversely correlated with progression-free survival in pancreatic cancer patients.
- Targeting IL-17B-IL-17RB signaling with an anti-IL-17RB antibody blocks pancreatic cancer metastasis by silencing multiple chemokines, HH Wu, WW Hwang-Verslues, WH Lee, CK Huang, PC Wei, CL Chen, JY Shew, EY Lee, YM Jeng, YW Tien, C Ma, WH Lee*, J Exp Medicine, 212, 333-349 (2015).
- Structural basis of interleukin-17B receptor in complex with a neutralizing antibody for guiding humanization and affinity maturation. WH Lee, X Chen, IJ Liu, JH Lee, CM Hu, HC Wu, SK Wang, WH Lee* and C Ma*. Cell Reports 41(4):111555 (2022).
- In August 2022, the technologies related to COVID-19 vaccine and antibodies have been transferred to a new startup biotech company by Academia Sinica for developing products such as broad-spectrum vaccines and antibodies with glycoengineering technologies.
RESEARCH HIGHLIGHTS:
- 2023/04/07, Structure of the heterotrimeric membrane protein complex FtsB-FtsL-FtsQ of the bacterial divisome
- 2023/01/30, Structural basis for a conserved neutralization epitope on the receptor-binding domain of SARS-CoV-2
- 2022/10/26, Crystal structure of 17B receptor complexed with antibody D9 provides solution for guiding antibody humanization and affinity maturation
- 2022/03/03, Mono-GlcNAc-decorated spike vaccine is highly effective against COVID-19 variants
- 2022/02/15, mRNA vaccine of SARS-CoV-2 spike protein with deletion of multiple glycosites is broadly protective against variants of concern
- 2021/01/27, Repurposing Existing Medicines as Anti-COVID-19 Virus Agents
- 2020/08/11, Monoglycosylated Chimeric Hemagglutinin as Universal Flu Vaccine
- 2020/07/29, FRIL Can Grab the Thorny Crown to Fight Off COVID-19
- 2019/02/20, Hassle Free Influenza Vaccine Close to Reality
- 2014/01/23, New Development of Universal Flu Vaccine
- 2009/10/15, Congratulations to Dr. Che Ma for being selected as the Young Affiliates by TWAS
- 2009/10/11, Removing the Sugar Coat of Influenza Hemagglutinin Proved Better Strategy In Vaccine Design
- 2009/08/25, PBP1b Research Rippled
- 2009/05/21, Key Structure of Membrane Protein Unveiled For Finding Next Antibiotics
- 2008/01/14, Discovery Of Antibiotics to Tackle the Problem of Drug Resistance
SELECTED PUBLICATIONS:
- Kung CC, Lo JM., Liao KS, Wu CY, Cheng LC, Chung C, Hsu TL, Ma C*, Wong CH*, 2025, “Expression of Human β3GalT5–1 in Insect Cells as Active Glycoforms for the Efficient Synthesis of Cancer-Associated Globo-Series Glycans”, Journal of the American Chemical Society, 147(13), 10864-10874. (SCIE) (IF: 15; SCI ranking: 9.6%)
- Lo JM., Kung CC, Cheng TJR, Wong CH*, Ma C*, 2025, “Structure-Based Mechanism and Specificity of Human Galactosyltransferase β3GalT5”, Journal of the American Chemical Society, 147(13), 10875-10885. (SCIE) (IF: 15; SCI ranking: 9.6%)
- Chen X, Mohapatra A, Nguyen HTV, Schimanski L, Kit TT, Rijal P, Chen CP, Cheng SH, Lee WH, Chou YC, Townsend AR, Ma C, Huang KYA, 2024, “The presence of broadly neutralizing anti-SARS-CoV-2 RBD antibodies elicited by primary series and booster dose of COVID-19 vaccine”, PLOS Pathogens, 20(6), e1012246. (SCIE)
- Huang KA*, Chen X, Mohapatra A, Nguyen HTV, Schimanski L, Tan TK, Rijal P, Vester SK, Hills RA, Howarth M, Keeffe JR, Cohen AA, Kakutani LM, Wu YM, Shahed-Al-Mahmud M, Chou YC, Bjorkman PJ, Townsend AR, Ma C*, 2023, “Structural basis for a conserved neutralization epitope on the receptor-binding domain of SARS-CoV-2.”, Nature communications, 14(1), 311. (SCIE)
- Nguyen HTV, Chen X, Parada C, Luo AC, Shih O, Jeng US, Huang CY, Shih YL*, Ma C*, 2023, “Structure of the heterotrimeric membrane protein complex FtsB-FtsL-FtsQ of the bacterial divisome”, Nature Communications, 14(1), 1903. (SCIE)
- T Yong, KK Chang, YS Wang and C Ma*, 2022, “Active humoral response reverts tumorigenicity through disruption of key signaling pathway”, Vaccines, 10, 163. (SCIE)
- Daniell H, Nair SK, Guan H, Guo Y, Kulchar RJ, Torres MDT, Shahed-Al-Mahmud M, Wakade G, Liu YM, Marques AD, Graham-Wooten J, Zhou W, Wang P, Molugu SK, de Araujo WR, de la Fuente-Nunez C, Ma C, Short WR, Tebas P, Margulies KB, Bushman FD, Mante FK, Ricciardi RP, Collman RG, Wolff MS, 2022, “Debulking different Corona (SARS-CoV-2 delta, omicron, OC43) and Influenza (H1N1, H3N2) virus strains by plant viral trap proteins in chewing gums to decrease infection and transmission.”, Biomaterials, 288, 121671. (SCIE)
- Wu CY, Cheng CW, Kung CC, Liao KS, Jan JT, Ma C, Wong CH*, 2022, “Glycosite-deleted mRNA of SARS-CoV-2 spike protein as a broad-spectrum vaccine.”, Proceedings of the National Academy of Sciences of the United States of America, 119(9), e2119995119. (SCIE)
- Lee WH, Chen X, Liu IJ, Lee JH, Hu CM, Wu HC, Wang SK, Lee WH, Ma C*, 2022, “Structural basis of interleukin-17B receptor in complex with a neutralizing antibody for guiding humanization and affinity maturation”, Cell Reports, 41(4), 111555. (SCIE)
- KYA Huang, D Zhou, TK Tan, C Chen, HME Duyvesteyn, Y Zhao, HM Ginn, L Qin, P Rijal, L Schimanski, R Donat, A Harding, J Gilbert-Jaramillo, W James, JA Tree, K Buttigieg, M Carroll, S Charlton, CE Lien, MY Lin, CP Chen, SH Cheng, X Chen, TY Lin, EE Fry, J Ren, C Ma, AR Townsend, DI Stuart, 2022, “Structures and therapeutic potential of anti-RBD human monoclonal antibodies against SARS-CoV-2”, Theranostics, 12, 1-17. (SCIE)
- HY Huang, HY Liao, X Chen, SW Wang, CW Cheng, M Shahed-Al-Mahmud, YM Liu, A Mohapatra, TH Chen, JM Lo, YM Wu, HH Ma, YH Chang, HY Tsai, YC Chou, YP Hsueh, CY Tsai, PY Huang, SY Chang, TL Chao, HC Kao, YM Tsai, YH Chen, CY Wu, JT Jan, TJR Cheng, KI Lin*, C Ma* and CH Wong*, 2022, “Vaccination with SARS-CoV-2 spike protein lacking glycan shields elicits enhanced protective responses in animal models”, Science Translational Medicine, 14, eabm0899. (SCIE)
- Ko YA, Yu YH, Wu YF, Tseng YC, Chen CL, Goh KS, Liao HY, Chen TH, Cheng TR, Yang AS, Wong CH, Ma C, Lin KI, 2021, “A non-neutralizing antibody broadly protects against influenza virus infection by engaging effector cells.”, PLoS pathogens, 17(8), e1009724. (SCIE)
- Huang KYA, Tan TK, Chen TH, Huang CG, Harvey R, Hussain S, Chen CP, Harding A, Gilbert-Jaramillo J, Liu X, Knight M, Schimanski L, Shih SR, Lin YC, Cheng CY, Cheng SH, Huang YC, Lin TY, Jan JT, Ma C, James W, Daniels RS, McCauley JW, Rijal P, Townsend AR, 2021, “Breadth and function of antibody response to acute SARS-CoV-2 infection in humans”, PLOS Pathogens, 17(2), e1009352. (SCIE)
- Jan JT, Cheng TR, Juang YP, Ma HH, Wu YT, Yang WB, Cheng CW, Chen X, Chou TH, Shie JJ, Cheng WC, Chein RJ, Mao SS, Liang PH*, Ma C*, Hung SC*, Wong CH*, 2021, “Identification of existing pharmaceuticals and herbal medicines as inhibitors of SARS-CoV-2 infection.”, Proceedings of the National Academy of Sciences of the United States of America, 118(5). (SCIE)
- Liu YM, Shahed-Al-Mahmud M, Chen X, Chen TH, Liao KS, Lo JM, Wu YM, Ho MC, Wu CY, Wong CH, Jan JT, Ma C*, 2020, “A carbohydrate-binding protein from the edible Lablab beans effectively blocks the infections of influenza viruses and SARS-CoV-2”, Cell Reports, 32, 108016. (SCIE)
- Chen JR, Liu YM, Tseng YC, Ma C*, 2020, “Better influenza vaccines: an industry perspective”, Journal of Biomedical Science, 27(1), 33. (SCIE)
- Liao HY, Wang SC, Ko YA, Lin KI, Ma C, Cheng TR, Wong CH, 2020, “Chimeric hemagglutinin vaccine elicits broadly protective CD4 and CD8 T cell responses against multiple influenza strains and subtypes.”, Proceedings of the National Academy of Sciences of the United States of America, 117(30), 17757-17763. (SCIE)
- Zhou D, Duyvesteyn HME, Chen CP, Huang CG, Chen TH, Shih SR, Lin YC, Cheng CY, Cheng SH, Huang YC, Lin TY, Ma C, Huo J, Carrique L, Malinauskas T, Ruza RR, Shah PNM, Tan TK, Rijal P, Donat RF, Godwin K, Buttigieg KR, Tree JA, Radecke J, Paterson NG, Supasa P, Mongkolsapaya J, Screaton GR, Carroll MW, Gilbert-Jaramillo J, Knight ML, James W, Owens RJ, Naismith JH, Townsend AR, Fry EE, Zhao Y, Ren J, Stuart DI, Huang KYA, 2020, “Structural basis for the neutralization of SARS-CoV-2 by an antibody from a convalescent patient”, Nature Structural & Molecular Biology, 27, 950. (SCIE)
- Huang CF, Chang WH, Lee TK, Joti Y, Nishino Y, Kimura T, Suzuki A, Bessho Y, Lee TT, Chen MC, Yang SM, Hwu Y, Huang SH, Li PN, Chen P, Tseng YC, Ma C, Hsu TL, Wong CH, Tono K, Ishikawa T, Liang KS, 2020, “XFEL coherent diffraction imaging for weakly scattering particles using heterodyne interference”, AIP Advances, 10(5), 055219. (SCIE)
- Tseng YC, Wu CY, Liu ML, Chen TH, Chiang WL, Yu YH, Jan JT, Lin KI, Wong CH*, and Ma C*, 2019, “Egg-based influenza split virus vaccine with monoglycosylation induces cross-strain protection against influenza virus infections”, PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA, 116(10), 4200-4205. (SCIE)
- Chen X, Wong CH, Ma C*, 2019, “Targeting the Bacterial Transglycosylase: Antibiotic Development from a Structural Perspective”, ACS Infectious Diseases, 5(9), 1493-1504. (SCIE)
- Huang KA, Rijal P, Jiang H, Wang B, Schimanski L, Dong T, Liu YM, Chang P, Iqbal M, Wang MC, Chen Z, Song R, Huang CC, Yang JH, Qi J, Lin TY, Li A, Powell TJ, Jan JT, Ma C, Gao GF, Shi Y, Townsend AR, 2018, “Structure-function analysis of neutralizing antibodies to H7N9 influenza from naturally infected humans.”, Nature microbiology, 4(2), 306-315. (SCIE)
- Chen CL, Hsu JC, Lin CW, Wang CH, Tsai MH, Wu CY, Wong CH*, Ma C*, 2017, “Crystal Structure of a Homogeneous IgG-Fc Glycoform with the N-Glycan Designed to Maximize the Antibody Dependent Cellular Cytotoxicity.”, ACS chemical biology, 12(5), 1335-1345. (SCIE)
- Chen CP, Lin MH, Chan YT, Chen LC, Ma C*, Fischer WB*, 2016, “Membrane protein assembly: two cytoplasmic phosphorylated serine sites of Vpu from HIV-1 affect oligomerization.”, Scientific reports, 6, 28866. (SCIE)
- Lin CW, Tsai MH, Li ST, Tsai TI, Chu KC, Liu YC, Lai MY, Wu CY, Tseng YC, Shivatare SS, Wang CH, Chao P, Wang SY, Shih HW, Zeng YF, You TH, Liao JY, Tu YC, Lin YS, Chuang HY, Chen CL, Tsai CS, Huang CC, Lin NH, Ma C, Wu CY, Wong CH, 2015, “A common glycan structure on immunoglobulin G for enhancement of effector functions.”, Proceedings of the National Academy of Sciences of the United States of America, 112(34), 10611-6. (SCIE)
- Wright JD., Chu HM, Huang CH, Ma C, Chang TW, Lim C, 2015, “Structural and Physical Basis for Anti-IgE Therapy”, Scientific Reports, 5(1), 11581. (SCIE)
- Wu HH, Hwang-Verslues WW, Lee WH, Huang CK, Wei PC, Chen CL, Shew JY, Lee EY, Jeng YM, Tien YW, Ma C, Lee WH, 2015, “Targeting IL-17B-IL-17RB signaling with an anti-IL-17RB antibody blocks pancreatic cancer metastasis by silencing multiple chemokines.”, The Journal of experimental medicine, 212, 333-349. (SCIE)
- Huang CK, Yang CY, Jeng YM, Chen CL, Wu HH, Chang YC, Ma C, Kuo WH, Chang KJ, Shew JY, Lee WH, 2014, “Autocrine/paracrine mechanism of interleukin-17B receptor promotes breast tumorigenesis through NF-kappaB-mediated antiapoptotic pathway.”, Oncogene, 33, 2968-2977. (SCIE)
- Chen PC, Chuang PK, Chen CH, Chan YT, Chen JR, Lin SW, Ma C, Hsu TL, Wong CH, 2014, “Role of N-linked glycans in the interactions of recombinant HCV envelope glycoproteins with cellular receptors.”, ACS chemical biology, 9(7), 1437-1443. (SCIE)
- Kung CC, Naik MT, Wang SH, Shih HM, Chang CC, Lin LY, Chen CL, Ma C, Chang CF, Huang TH, 2014, “Structural analysis of poly-SUMO chain recognition by the RNF4-SIMs domain.”, The Biochemical journal, 462(1), 53-65. (SCIE)
- Chen JR, Yu YH, Tseng YC, Chiang WL, Chiang MF, Ko YA, Chiu YK, Ma HH, Wu CY, Jan JT, Lin KI*, Ma C*, Wong CH*, 2014, “Vaccination of monoglycosylated hemagglutinin induces cross-strain protection against influenza virus infections.”, Proceedings of the National Academy of Sciences of the United States of America, 111(7), 2476-2481. (SCIE)
- Huang CY, Shih HW, Lin LY, Tien YW, Cheng TJR, Cheng WC, Wong CH*, and Ma C*, 2012, “Crystal structure of Staphylococcus aureus transglycosylase in complex with a lipid II analog and elucidation of peptidoglycan synthesis mechanism”, Proceedings of the National Academy of Sciences of the United States of America, 109(17), 6496-6501. (SCIE)
- Shih HW, Chang YF, Li WJ, Meng FC, Huang CY, Ma C, Cheng TJ, Wong CH, Cheng WC, 2012, “Effect of the peptide moiety of Lipid II on bacterial transglycosylase.”, Angewandte Chemie-International Edition, 51(40), 10123-10126. (SCIE)
- Chen JR, Ma C*, Wong CH*, 2011, “Vaccine design of hemagglutinin glycoprotein against influenza.”, Trends in biotechnology, 29, 426-434. (SCIE)
- Sung MT, Lai YT, Huang CY, Chou LY, Shih HW, Cheng WC, Wong CH* and Ma C*, 2009, “Crystal structure of the membrane-bound bifunctional transglycosylase PBP1b from Escherichia coli”, Proceedings of the National Academy of Sciences of the United States of America, 106, 8824-8829. (SCIE)
- Wang CC, Chen JR, Tseng YC, Hsu CH, Hung YF, Chen SW, Chen CM, Khoo KH, Cheng TJ, Cheng YSE, Jan JT, Wu CY, Ma C* and Wong CH*, 2009, “Glycans on influenza hemagglutinin affect receptor binding and immune response”, Proceedings of the National Academy of Sciences of the United States of America, 106, 18137-18142. (SCIE)
- Cheng TJR, Sung MT, Liao HY, Chang YF, Chen CW, Huang CY, Chou LY, Wu YD, Chen YH, Cheng YSE, Wong CH*, Ma C* and Cheng WC*, 2008, “Domain requirement of moenomycin binding to bifunctional transglycosylases and development of high-throughput discovery of antibiotics”, Proceedings of the National Academy of Sciences of the United States of America, 105, 431-436. (SCIE)