 副研究員
副研究員
02-27871246
Lab phone: 02-27898804
EDUCATION AND POSITIONS HELD:
- Ph.D., Environmental Toxicology, University of California-Riverside, June 2007
- Master of Environmental Management, Environmental Toxicology, Chemistry and Risk Assessment, Duke University, May 2002
- Bachelor of Science, Geography, National Taiwan University, Taipei, Taiwan, June 1999
- Instructor, Department of Public Health, Loma Linda University, Loma Linda, CA, U.S.A., 2006, 2007
- Postdoctoral Fellow, Genomics Research Center, Academia Sinica, Jan 2008- June 2014
- Research Scientist, Department of Biology, Washington University in St. Louis, Sep 2014- May 2015
- Assistant Professor, Genomics Research Center, Academia Sinica, Dec 2015- June 2023
- Associate Professor, Genomics Research Center, Academia Sinica, Taiwan, June 2023-present
HONORS:
- Taiwan National Science Council 2013 Postdoctoral Fellow Academic Publication Award
- The 72nd Annual Meeting of the Japanese Cancer Association Travel Grant Award Oct 2013
- The 1st GRC Best Performance-and-Service Award, Genomics Research Center, Academia Sinica, Feb 2012
- The 70th Annual Meeting of the Japanese Cancer Association Travel Grant Award Oct 2011
- Academia Sinica Postdoctoral Research Fellowship Jan 2010- Dec 2011
- Academia Sinica Distinguished Postdoctoral Scholar Fellowship Jan 2008- Dec 2009
- Sigma Xi Grants-in-Aid of Research Award, Jan 2007
- Fukuto Award, University of California-Riverside June 2006
RESEARCH INTERESTS:
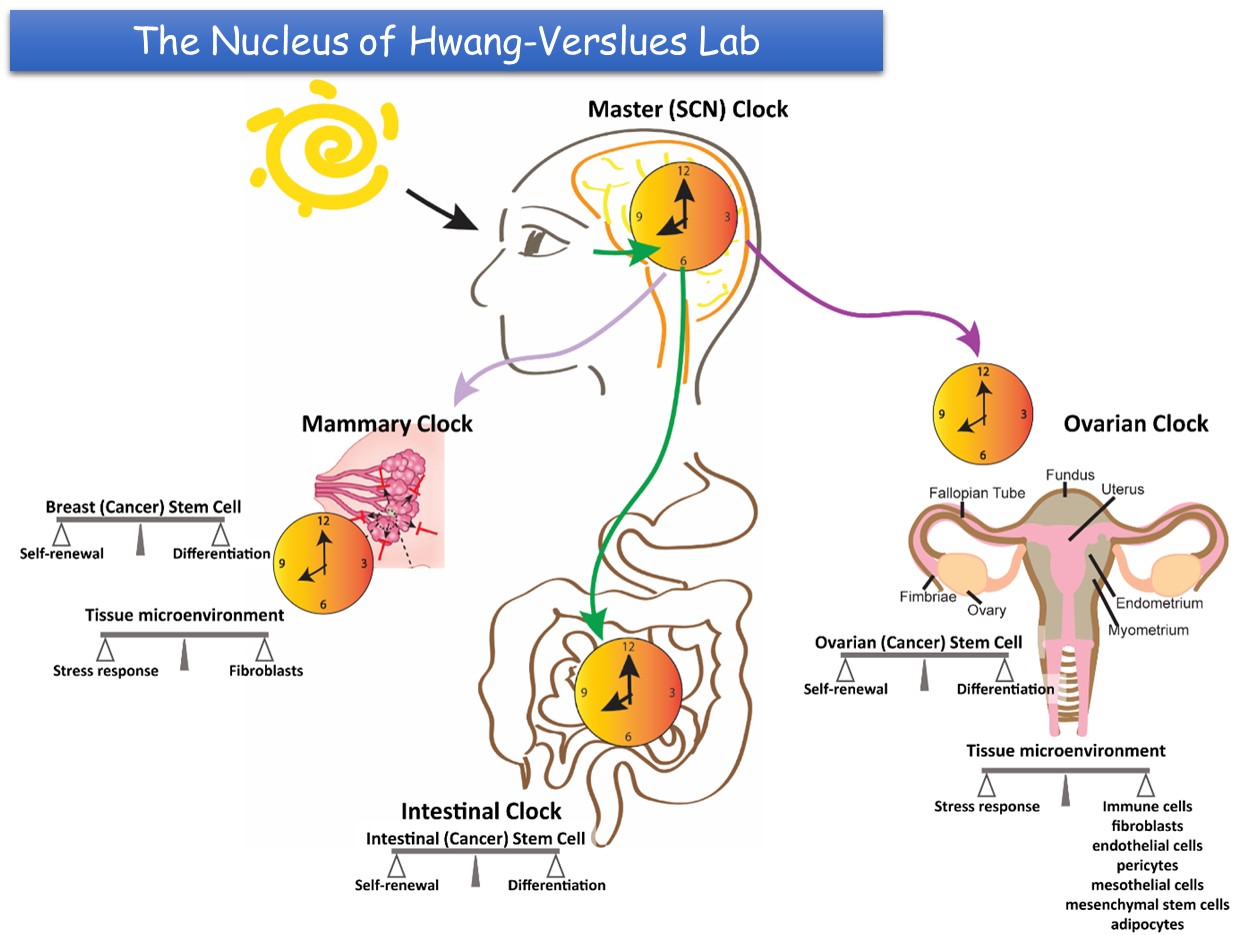
The main focus of the Hwang-Verslues laboratory is on cancer and circadian biology. We have been pursuing three research aims: 1. Identify key factors critical for cancer initiation, metastasis and recurrence, 2. Determine the significance of circadian and core clock gene regulation in tissue homeostasis and stress response, and 3. Investigate the interplay between circadian dysregulation and cancer progression. For each of these research aims, our accomplishments to date are summarized as below.
Research aim 1: Key factors driving tumorigenesis and cancer metastasis and recurrence
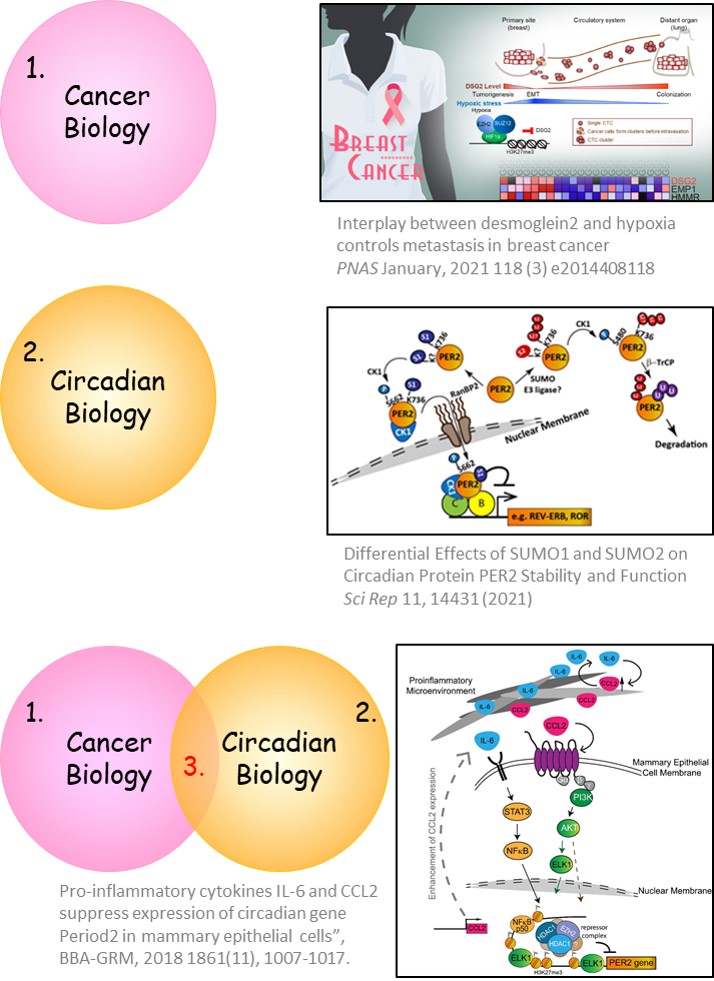
Breast cancer: Interplay between desmoglein2 and hypoxia controls metastasis in breast cancer [Chang PH et al, PNAS, 2021 (PMID: 33431674)]. We found that high DSG2 expression is correlated with poor prognosis and recurrence risk in breast cancer patients. DSG2 promotes tumor growth, increases CTC clusters and facilitates distant organ colonization. Dynamic repression and de-repression of DSG2 in response to changing levels of oxygen is needed to allow cells to detach from hypoxic primary tumor, invade in circulation and later colonize in oxygen rich distal organs. Also, this is the first report demonstrating a transcriptional suppression function of HIF-1α on an oncogenic factor. These findings demonstrate the importance of DSG2 in all stages of breast cancer metastasis. [supported by Academia Sinica (AS-SUMMIT-108, -109) and the Ministry of Science and Technology (MOST 105-2628-B-001-008-MY3)].
Ovarian cancer: The dual-function RNA binding protein MEX3A mediates p53 degradation to prevent ferroptosis and facilitate ovarian cancer tumorigenesis [Wang CK et al., in revision]. We found that MEX3A, which is a dual-function protein containing a RING finger domain and an RNA binding domain, is correlated with poor survival in OC. Further investigation revealed that MEX3A upregulation is crucial for tumorigenesis in ovarian cancer (OC) expressing wild-type (WT) p53 protein [e.g., ovarian clear cell carcinoma (OCCC) and endometrioid carcinoma (EC)]. MEX3A overexpression enhances tumorigenic activity; whereas, depletion of MEX3A reduces cell survival and proliferative ability, inhibits tumor growth and prolongs survival. MEX3A depletion does not alter p53 mRNA level but does increase the protein stability of WT p53. MEX3A-mediated p53 protein degradation is crucial to prevent ferroptosis and enhance tumorigenesis. Identification of MEX3A-WT p53-ferroptosis as a pathway determining clinical severity of ovarian tumorigenesis provides an opportunity to develop new treatment strategies for cases of OC with WT p53. [supported by Academia Sinica (AS-SUMMIT-108, -109) and the Ministry of Science and Technology (MOST111-2314-B-001-012-)].
We are currently investigating additional MEX3A functions in OC metastasis.
Research Aim 2: Circadian and core clock gene regulation in tissue homeostasis and stress response
Core circadian transcription factor BMAL1 and RNA-binding protein MEX3A co-regulate intestinal stem cell homeostasis and stress response. (Cheng LT and Chang FP et al. This study is currently under preparation for submission. bioRxiv doi: https://doi.org/10.1101/2022.03.08.483537). We found that the core clock gene BMAL1 participates in regulating intestinal stem cell (ISC) homeostasis. BMAL1 transcriptionally regulates oscillating expression of two essential ISC genes, Mex3a and Lgr5, in the crypt-based ISCs. Depletion of BMAL1 in the Lgr5+ ISCs perturbed lineage specific gene expression in the intestinal epithelium. BMAL1 appears to be essential for the intestinal epithelium to cope with stress. 5-FU delivery at the time of day when Lgr5 expression peaked increased apoptosis and inhibited intestinal epithelial cell proliferation. In contrast, apoptosis was not detectable if 5-FU was delivered when Lgr5 expression was lowest. These results provide mechanistic explanation of how chronotherapy can reduce 5-FU side effects. [supported by Academia Sinica (AS-SUMMIT-108, -109)]
We are currently investigating how BMAL1 participates in intestinal stress response.
Research aim 3: Interplay between circadian and cancer
PER2 suppression enhances tumorigenic potential and EMT in breast cancer. I previously discovered that core clock gene PER2 inhibits invasion and cancer stemness in breast cells [Hwang-Verslues et al., PNAS, 2013 (PMID: 23836662)]. Recently, we found that IL-6 and CCL2 in the inflammatory microenvironment suppressed expression of PER2 to increase tumorigenic potential in breast epithelial cells. These data show a new mechanism by which mammary cells interact with a cancerous microenvironment via PER2 [Yu CW et al, BBA-GRM, 2018 (PMID: 30343691)]. We also found that PER2 can be SUMOylated to modulate its transcriptional repressor function and protein stability. Thus, PER2 SUMOylation is also likely to affect the tumor suppressor functions of PER2 as well as PER2 clock function. [Chen et al, Scientific Reports, 2021 (PMID:342573720)].
We are currently investigating BMAL and PER2 functions in ovarian cancer.

SELECTED PUBLICATIONS:
- Cheng L.-T., Tan G.Y.T., Chang F.-P., Wang C.-K., Chou Y.-C., Hsu P.-H., Hwang-Verslues W.W.*, 2023, “Core clock gene BMAL1 and RNA-binding protein MEX3A collaboratively regulate Lgr5 expression in intestinal crypt cells”, SCIENTIFIC REPORTS, 13, 17597. (SCIE)
- Wang C.-K., Chen T.-J., Tan G.Y.T., Chang F.-P., Sridharan S., Yu C.-H. A., Chang Y.-H., Chen Y.-J., Cheng L.-T., Hwang-Verslues W.W.*, 2023, “MEX3A mediates p53 degradation to suppress ferroptosis and facilitate ovarian cancer tumorigenesis”, Cancer Research, 83 (2), 251-263. (SCIE)
- Chen L.C., Hsieh Y.L., Tan G.Y.T., Kuo T.Y., Chou Y.C., Hsu P.H., Hwang-Verslues W.W.*, 2021, “Differential Effects of SUMO1 and SUMO2 on Circadian Protein PER2 Stability and Function”, SCIENTIFIC REPORTS, 11, 14431. (SCIE)
- Chang P.H., Chen M.C., Tsai Y.P., Tan G.Y.T., Hsu P.H., Jeng Y.M., Tsai Y.F., Yang M.H., Hwang-Verslues W.W.*, 2021, “Interplay between Desmoglein2 and hypoxia controls metastasis in breast cancer”, PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES OF THE UNITED STATES OF AMERICA, 118(3), e2014408118. (SCIE)
- Fararjeh, A., Tu, S.H., Chen, L.C., Cheng, T.C., Liu, Y.R., Chang, H.L., Chang, H.W., Huang, C.C., Wang, H.C.R., Hwang-Verslues, W.W., Wu, C.H., Ho, Y.S.*, 2019, “Long-term exposure to extremely low-dose of nicotine and 4-(methylnitrosamino)-1-(3-pyridyl)-1-butanone (NNK) induce non-malignant breast epithelial cell transformation through activation of the a9-nicotinic acetylcholine receptor-mediated signaling pathway”, ENVIRONMENTAL TOXICOLOGY, 34(1), 73-82. (SCIE)
- Yu, C.-W., Cheng, K.-C., Chen, L.-C., Lin, M.-X., Chang, Y.-C., Hwang-Verslues, W.W.*, 2018, “Pro-inflammatory cytokines IL-6 and CCL2 suppress expression of circadian gene Period2 in mammary epithelial cells”, BIOCHIMICA ET BIOPHYSICA ACTA-GENE REGULATORY MECHANISMS, 1861(11), 1007-1017. (SCIE)
- Abdul F.F.-S., Tu S.-H., Chen L.-C., Liu Y.-R., Lin Y.-K., Chang H.-L., Chang H.-W., Wu C.-H., Hwang-Verslues W.W., Ho Y.-S., 2018, “The Impact of the effectiveness of GATA3 as a prognostic factor in breast cancer”, Human Pathology, 80, 219-230. (SCIE)
- Wu YW, Hsu KC, Lee HY, Huang TC, Lin TE, Chen YL, Sung TY, Liou JP, Hwang-Verslues WW, Pan SL, Huang-Fu WC,, 2018, “A novel dual HDAC6 and tubulin inhibitor, MPT0B451, displays anti-tumor ability in human cancer cells in vitro and in vivo”, FRONTIERS IN PHARMACOLOGY, 9, 205. (SCIE)
- Huang S.-C., Wei P.-C., Hwang-Verslues W.W., Kuo W.-H., Jeng Y.-M., Hu C.-H., Shew J.-Y., Huang C.-S., Chang K.-J., Lee E. Y.-H. P., Lee W.-H.*, 2017, “TGF-β1 secreted by Tregs in lymph nodes promotes breast cancer malignancy via up-regulation of IL-17RB”, EMBO MOLECULAR MEDICINE, 9(12), 1660-1680. (SCIE)
- Cheung, S.K.C., Chuang, P.-K., Huang, H.-W., Hwang-Verslues, W.W., Cho C.H.-H., Yang, W.-B., Shen, C.-N., Hsiao, M., Hsu, T.-L., Chang, C.-F., Wong, C.-H., 2016, “Stage-Specific Embryonic Antigen-3 (SSEA-3) and β3GalT5 are Cancer Specific and Significant Markers for Breast Cancer Stem Cells”, Proceedings of the National Academy of Sciences of the United States of America, 113(4), 960-965. (SCIE)
- Wu, H.-H*, Hwang-Verslues, W.W.*, Lee, W.-H., Huang, C.-K., Wei, P.-C., Chen, C.-L. , Shew, J.-Y., Lee, E. Y.-H. P., Jeng, Y.-M., Tien, Y.-W., Ma, C., Lee, W.-H., 2015, “Targeting IL-17B/RB signaling with an anti-IL17RB antibody blocks pancreatic cancer metastasis by silencing multiple chemokines”, JOURNAL OF EXPERIMENTAL MEDICINE, 212(3), 333-349. (SCIE)
- Hwang-Verslues WW, Chang PH, Jeng YM, Kuo WH, Chiang PH, Chang YC, Hsieh TH, Su FY, Lin LC, Abbondante S, Yang CY, Hsu HM, Yu JC, Chang KJ, Shew JY, Lee EY, Lee WH, 2013, “Loss of corepressor PER2 under hypoxia up-regulates OCT1-mediated EMT gene expression and enhances tumor malignancy.”, Proceedings of the National Academy of Sciences of the United States of America, 110(30), 12331-6. (SCIE)
- Chang, P.-H., Hwang-Verslues, W.W., Chang, Y.-C., Chen, C.-C., Hsiao, M., Jeng, Y.-M., Chang, K.-J., Lee, E. Y.-H. P., Shew, J.-Y., Lee, W.-H., 2012, “Activation of Robo1 signaling of breast cancer cells by Slit2 from stromal fibroblast restrains tumorigenesis via blocking PI3K/Akt/β-catenin pathway”, CANCER RESEARCH, 72(18), 4652-4661. (SCIE)
- Hwang-Verslues, W.W.*, Lee, W.-H., Lee, E.Y.-H.P., 2012, “Biomarkers to target heterogeneous breast cancer stem cells (invited review)”, Journal of Molecular Biomarkers & Diagnosis, S8, 006.
- Hwang-Verslues, W.W., Chang P.-H., Wei P.-C., Yang C.-Y., Huang C.-K., Kuo W.-H., Shew J.-Y., Chang K.-J., Lee E.-Y., Lee W.-H., 2011, “miR-495 is upregulated by E12/E47 in breast cancer stem cells, and promotes oncogenesis and hypoxia resistance via downregulation of E-cadherin and REDD1”, ONCOGENE, 30(21), 2463-2474. (SCIE)
- Hwang-Verslues, W.W., Sladek, F.M., 2010, “HNF4a-role in drug metabolism and potential drug target? (review)”, CURRENT OPINION IN PHARMACOLOGY, 10(6), 698-705. (SCIE)
- Bolotin E., Liao H., Ta T.C., Yang C., Hwang-Verslues W.W., Evans J.R., Jiang T., Sladek F.M., 2010, “Integrated approach for identification of HNF4α target genes using protein binding microarrays”, HEPATOLOGY, 51(2), 642-653. (SCIE)
- Hwang-Verslues, W.W., Kuo W.-H., Chang P.-H. Pan C.-C., Wang H.-H., Tsai S.-T., Jeng Y.-M., Shew J.-Y., Kung J.T, Chen C.-H., Lee E. Y.-H. P., Chang K.-J., Lee W.-H., 2009, “Multiple lineages of human breast cancer stem/progenitor cells identified by profiling with stem cell markers”, PLoS One, 4(12), e8377. (SCIE)
- Hwang-Verslues, W.W., Chang, K.J., Lee, E.Y-H.P., Lee, W.-H., 2008, “Breast cancer stem cells and tumor suppressor genes”, Journal of Formosan Medical Association, 107(10), 751-766.
- Hwang-Verslues, W.W., Sladek, F.M., 2008, “Nuclear receptor HNF4alpha1 competes with oncoprotein c-Myc for control of the p21/WAF1 promoter”, MOLECULAR ENDOCRINOLOGY, 22(1), 78-90.
- Maeda, Y.*, Hwang-Verslues, W.W.*, Wei, G., Fukazawa, T., Durbin, M.L., Owen, L. B., Liu, X. and Sladek, F.M., 2006, “Tumor suppressor p53 down regulates the expression of the human hepatocyte nuclear factor 4alpha (HNF4a) gene”, BIOCHEMICAL JOURNAL, 400(2), 303-313. (SCIE)