Since the outbreak of severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) in December 2019 that caused Coronavirus Induced Disease 2019 (COVID-19), the virus has spread all over the world and caused more than 411 million infections and 5.8 million deaths in 25 months. Vaccination is an effective strategy to control the spread of SARS-CoV-2, and the protective antibodies induced by the vaccine are known to recognize the viral surface spike (S) protein and primed the T cell response. However, current vaccines may lose their efficacy because of the high mutation rate of SARS-CoV-2. To overcome this problem, development of a universal SARS-CoV-2 vaccine with broad protection against the current and upcoming variants is needed.
The team of Professor Che Ma, Kuo-I Lin and Chi-Huey Wong previously showed that immunization with mono-GlcNAc decorated S protein (SMG) induced broadly protective antibody and CD4+ as well as CD8+ T cell responses against SARS-CoV-2 and the variants of concern. In addition, using the single B cell technology to screen the B cells from SMG immunized mice, the team has identified a broadly neutralizing monoclonal antibody targeting the highly conserved epitope in the receptor binding domain (RBD) which was not induced in the fully glycosylated S immunized mice.
In this study, Dr. Chung-Yi Wu, a postdoctoral fellow, designed and studied the mRNA vaccines of SARS-CoV-2 spike protein with deletion of the glycosites in the RBD or the subunit 2 (S2) domain. The vaccines were found to elicit antibodies with 5-8 times more protection against the alpha, beta, gamma, delta and omicron variants, as compared to the unmodified mRNA (Fig.1). Alignment of the more than 6 million S protein sequences from GISAID by another postdoctoral fellow Dr. Cheng-Wei Cheng reveals that all the 24 glycosites in S protein are conserved and the 12 most conserved epitopes identified are located in the RBD and the S2 domains. In addition, deletion of all the 4 glycosites in RBD or especially all the 9 glycosites in S2 would expose the conserved epitopes to elicit broadly protective immune responses as these conserved epitopes are largely shielded by the glycans.
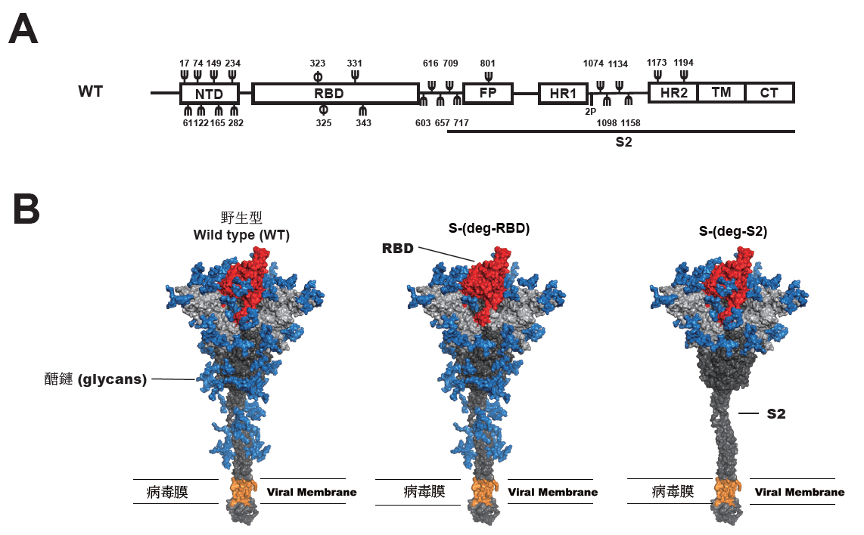 |
| Figure 1. A, Schematic representation of the SARS-CoV-2 spike protein. NTD, N-terminal domain. RBD, a receptor-binding domain. FP, the fusion peptide. HR1, heptapeptide repeat sequence 1. HR2, heptapeptide repeat sequence 2. TM, transmembrane domain. CT, cytoplasm domain. S2 subunit. 2P, (K986P, and V987P). Ψ, the N-glycosylation site; φ, O-glycosylation site. B, 3D image of the SARS-CoV-2 spike protein and vaccine design. Blue: glycan; Red: RBD; Gray: S1; Dark gray: S2; Orange: Transmembrane Domain. |
The study also found that the glycosylation on S protein affected protein folding and T cell response. Of particular significance in this study is that deletion of the glycosites in S2 led to the expression of misfolded S protein which was found to induce 2-3 times more CD8+ T cell immune response as compared to the unmodified mRNA. It enhanced the major histocompatibility complex class I expression in dendritic cells.
These findings suggest that removal of the glycans that shield the conserved epitopes can induce broader and stronger immune responses. This is the first glycosite-deleted SARS-CoV-2 mRNA vaccine developed which was shown to have a broad-spectrum of protection against SARS-CoV-2 and the five variants of concern, including the alpha, beta, gamma, delta and omicron variants, and this new strategy could be applied to the development of other mRNA vaccines.
It is difficult or impossible to express the improperly folded protein with deletion of certain glycosylation sites for immunization study. In this study, the team found that using mRNA as vaccine, they were able to address this problem and study the impact of protein glycosylation on immune response in vivo through expression of the de-glycosylated protein inside the cell. Therefore, this work represents a new milestone in vaccine development.
The first author of this paper is Dr. Chung-Yi Wu (a postdoctoral fellow at Genomics Research Center). Other authors include Dr. Cheng-Wei Cheng (currently an assistant professor at Kaohsiung Medical University), Mr. Chih-Chuan Kung, Dr. Kuo-Shiang Liao, Dr. Jia-Tsrong Jan, Dr. Che Ma and Dr. Chi-Huey Wong.
The complete result of this study can be read online at: C.-Y. Wu, C.-W. Cheng, C.-C. Kung, K.-S. Liao, J.-T. Jan, C. Ma, C.-H. Wong, “Glycosite-deleted mRNA of SARS-CoV-2 Spike Protein as Broad-Spectrum Vaccine” Proc. Natl. Acad. Sci. USA, 2022, doi:10.1073/pnas.2119995119.
This work was supported by the funding from Academia Sinica and from The Ministry of Science and Technology Translational Medical Research Program (MOST 109-0210-01-18-02)