Coronavirus disease 2019 (COVID-19) caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has resulted in more than 440 million confirmed cases and over 6 million deaths so far. Although there are COVID-19 vaccines available, viruses constantly change through mutations. Key questions remain about how quickly the variants can spread, their potential to evade immunity and how effective the vaccines are against the variants.
A research group, led by Dr. Chi-Huey Wong and Dr. Alex Ma from the Genomics Research Center, recently developed a Mono-GlcNAc-decorated spike protein vaccine, designed to protect against many different variants of coronaviruses. In the animal experiments, this universal coronavirus vaccine elicits a stronger immune response and provide better protection against the reported virus variants, including the, the UK Alpha variant , the Brazil Gamma variant and the Indian Delta variant.
More than one million S protein sequences with over 1,000 sites of mutation in its 1,273 amino acids have been reported to the GISAID database, including the highly transmissible Delta variant strains. The new mutations may decrease the ability of antibodies to neutralize the virus, and become a major challenge in the development of broadly protective vaccines to control the pandemic.
From the past experience of working with influenza virus, Dr. Wong and Dr. Ma’s team has studied the impact of glycosylation on the interaction with ACE2 receptor and spike glycoprotein expressed in different cell lines. The trimeric spike glycoprotein of SARS-CoV-2 is heavily glycosylated, influencing protein folding and evading host immune response. Removal of the S protein glycan shield may expose more highly conserved regions for eliciting neutralizing antibodies in a more efficient way.
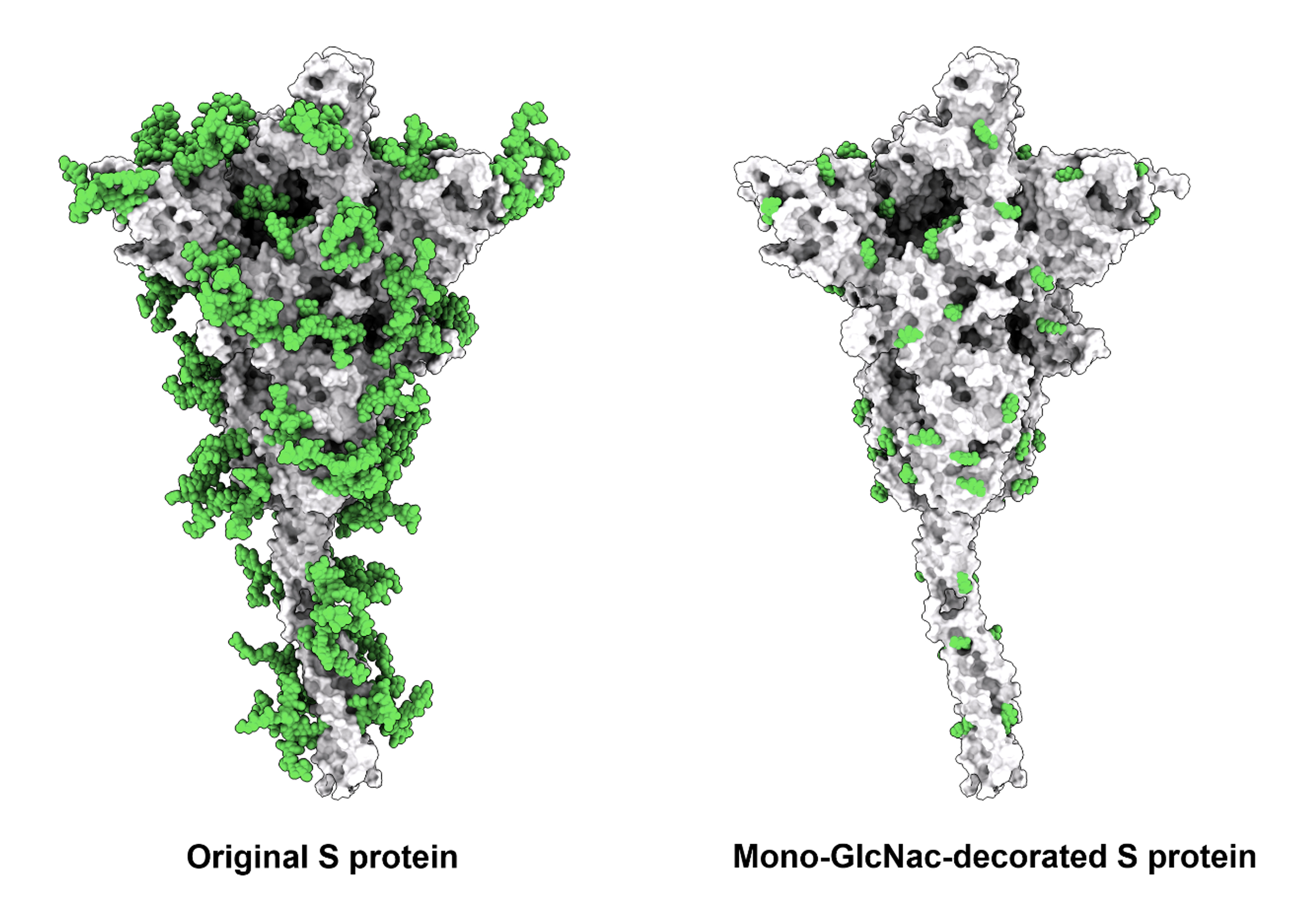
The team came up with a solution to cope with virus variants. “Removal of unnecessary glycans from S protein to better expose the highly conserved sequences is an effective approach to developing broadly protective vaccines against SARS-CoV-2 and variants.” Dr. Ma said.
Based on this approach, the research group has decided to design new spike glycoprotein vaccines through glycoengineering, by successfully removing the glycans that may interfere with the presentation of conserved epitopes and generating the Mono-GlcNAc-decorated form of S protein (SMG) which was further confirmed to be near 100% pure with single sugar residue at every N-glycosylation site.
In an effort to develop universal vaccines, the researchers found that mice immunized with Mono-GlcNAc-decorated S protein (SMG) elicited better antibody responses capable of neutralizing not only the wild type but also the variants of concerns including Alpha, Beta, Gamma and Delta variants than those with the fully-glycosylated S protein (SFG), and SMG vaccination provided 80% survival for hACE2 transgenic mice when challenged with lethal dose of SARS-CoV-2 Delta variant when SFG¬ vaccine only showed 20% survival. What's more, the vaccine spurred 3 folds more neutralizing antibodies against Delta than current vaccines do.
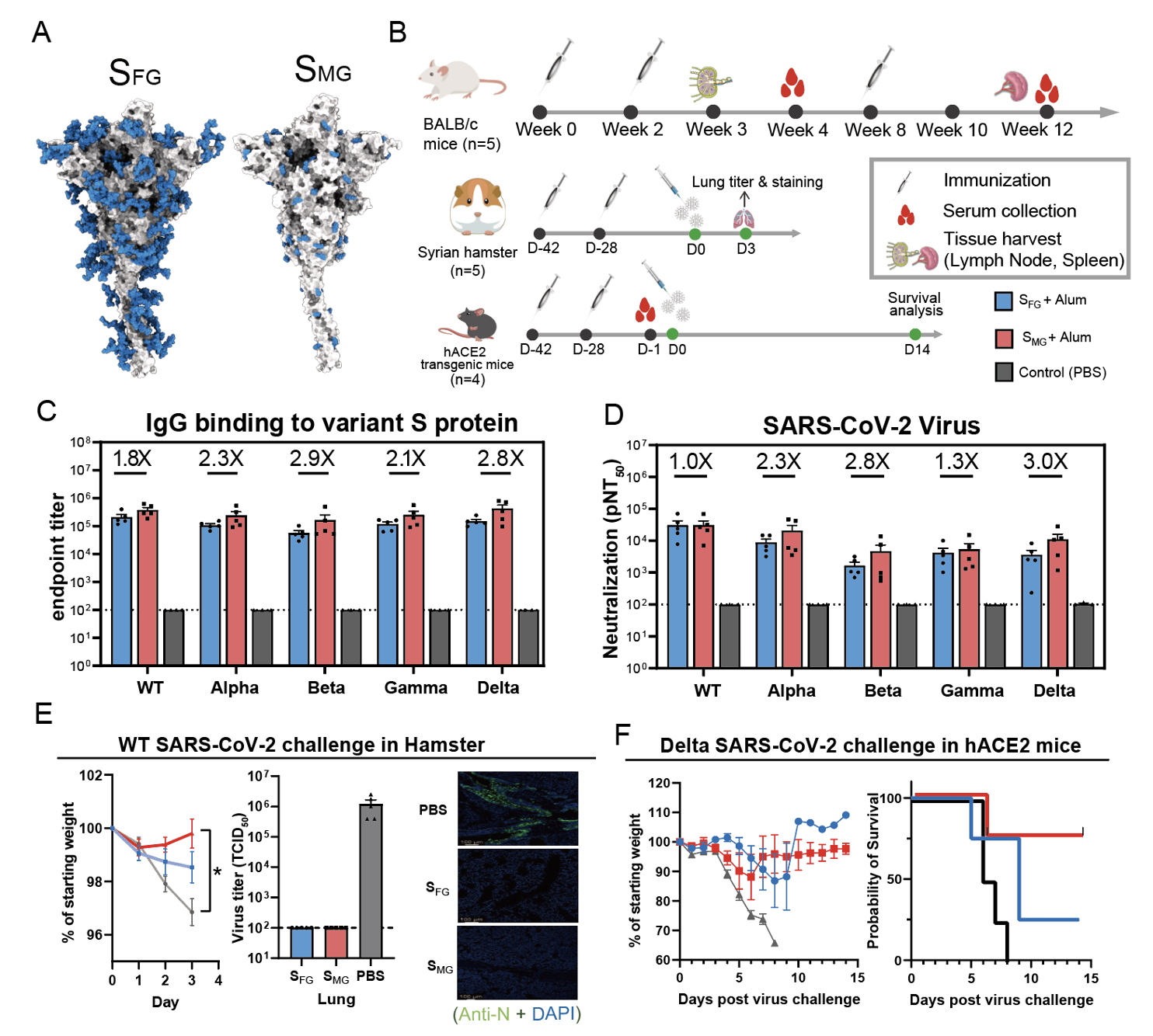
In addition, with Dr. Kuo-I Lin’s expertise, using single B cell technology, they isolated a monoclonal antibody from SMG immunized mice which was also able to neutralize the wild type and variants.
Like all viruses, SARS-CoV-2 continues to mutate as it spreads, and very likely will become a part of our life just like influenza. Developing a universal coronavirus vaccine is urgently needed to prevent possible future infections of SARS-CoV-2 variants with high transmissibility and mortality.
This research work was successfully completed by efforts from multiple GRC labs, including labs of Drs. Chi-Huey Wong, Alex Ma, Kuo-I Lin, Ting-Jen Rachel Cheng, Jia-Tsrong Jan. Appreciations to outside institutes include: Drs. Yi-Ping Hsueh, Ching-Yen Tsai, from Institute of Molecular Biology; Dr. Yen-Hui Chen, from Institute of Biomedical Sciences; Dr. Yu-Chi Chou from Biomedical translational research center RNAi core; and Dr. Sui-Yuan Chang from NTUH. Han-Yi Huang, Dr. Hsin-Yu Liao, Dr. Xiaorui Chen and Dr. Szu-Wen Wang are the core team members and will continue the work.
This study has been published in the Science Translational Medicine on 1st March, 2022 and can be read online at: https://www.science.org/doi/10.1126/scitranslmed.abm0899
Aademia Sinica COVID-19 Related Technologies: https://iptt.sinica.edu.tw/shares/1117